Overview
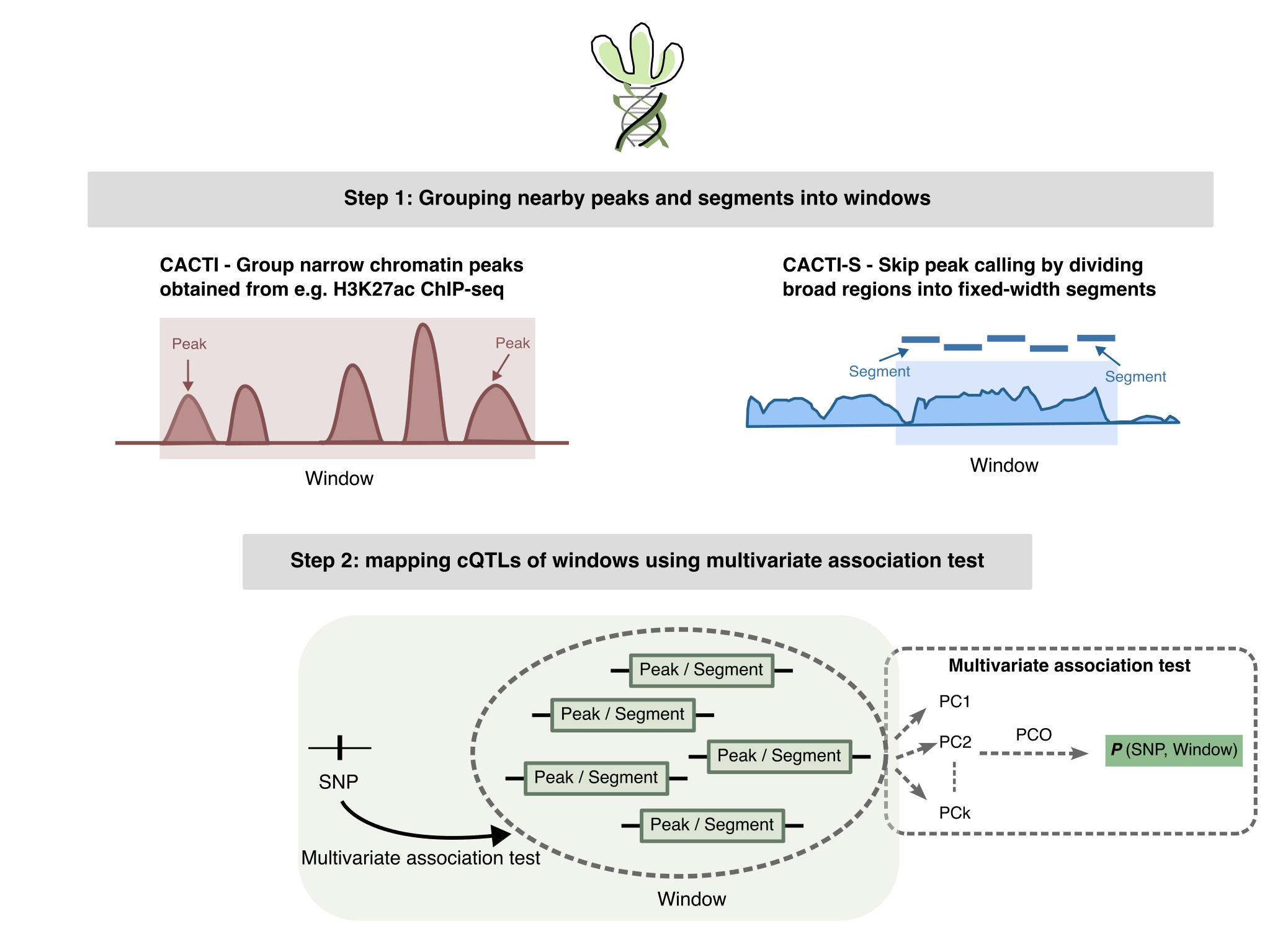
CACTI (ChromAtin quantitative loCi of mulTIple peaks) implements a powerful method for chromatin QTL mapping that leverages the correlation structure of nearby regulatory elements.
The package offers two main modules:
- CACTI (Peak-window method):
- Groups pre-defined peaks into non-overlapping windows of fixed genomic size.
- Residualizes peak intensities with respect to covariates.
- Runs multi-peak association tests using a principal-component omnibus (PCO) test.
- Aggregates results and computes window-level FDR.
- CACTI-S (Sample-based pipeline):
- An end-to-end preprocessing workflow to go from raw BAM files to normalized phenotype matrices to QTL mapping.
- Performs segmentation, read counting, QC/filtering, and normalization.
- Runs cis-QTL mapping to generate input files for CACTI.
Installation
install.packages("devtools") # if not installed yet
devtools::install_github("liliw-w/cacti", build_vignettes = TRUE)After installation:
Documentation
Vignettes
See the full documentation and vignettes for -
CACTI Peak-Window Pipeline
CACTI-S Pipeline
Vignettes can also be viewed within the installed package -